Phage contaminations threaten fermentation processes based on bacterial activities and pose a serious risk to companies relying on such bioprocesses, occasionally resulting in a complete paralysis of facility’s productivity. Although even the best practices do not guarantee contamination-free production, better prepared facility will recover faster, and the contamination will not turn into an outbreak.
PHAGE CONSULTANTS specializes in prevention and troubleshooting of phage contamination problems in various production and laboratory setups through a variety of services. We offer phage detection and identification services, facility audits and initial assessments, consultations, personnel training, advice on bioprocess optimization and SOP improvement, and new facility development. We also offer fast and reliable routine or as-required testing of incoming bacterial strains and cell banks for the presence of both lytic phages and prophages – a necessary tool in phage prevention. With these services, Phage Consultants can help you avoid phage contaminations and recover quickly from contaminations that have already occurred.
Our team of expert scientists working in state-of-the-art laboratories are ready to solve all bacteriophage-related problems. Not only can we help with phage contaminations, but we can also offer assistance with all aspects of bacteriophage use in various fields of biotechnology, medicine, food and crop protection etc., as well as contract phage manufacturing and phage purification services. Our extensive expertise in phage biology research helps us deliver high quality results in relatively short time and at competitive prices.
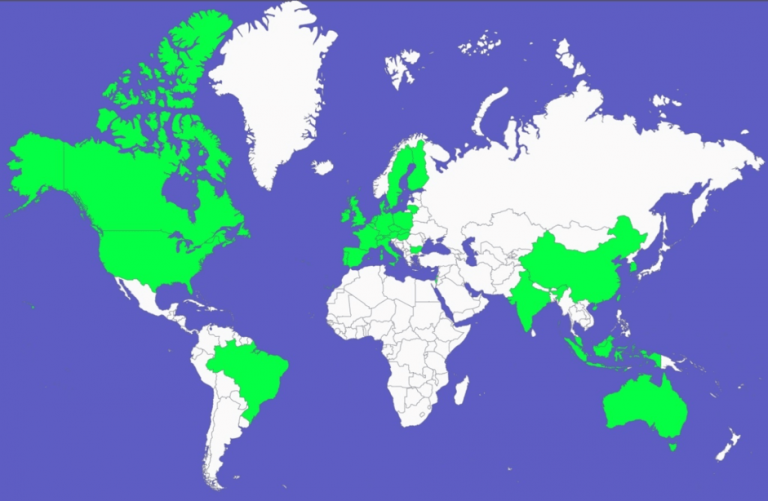
Since its establishment in 2007, Phage Consultants has successfully completed over 150 phage prevention and troubleshooting jobs worldwide (see the map below), and remains a global leader in prevention and resolution of phage-related production irregularities.
Phage Consultants is the category winner in „Microenterpreneur of the year” (2012) contest organized by CitiBank Foundation.
Phage Consultants received a third place award in 2012 edition of “Parkowe Orły” (“Park Eagles”) organized by InnoFirma and awarding companies with the highest innovative potential and with a presence on international market.
6. Project no. FEPM.01.01-IP.03-001/25 - "Opracowanie platformy PhageSense do wykrywania infekcji fagowych, stanowiącej innowację w obszarze nowoczesnych rozwiązań w diagnostyce". (Development of the PhageSense platform for detecting phage infections, representing an innovation in the field of modern diagnostic solutions).
Our publications from the field of phage-related research are listed below. Please use the “Contact Us” form to request a copy of any of the publications.
- Raza, S., Bończak, B., Atamas, N., Karpińska, A., Ratajczyk, T., Łoś, M., Hołyst, R., & Paczesny, J. (2025). The activity of indigo carmine against bacteriophages: an edible antiphage agent. Applied Microbiology and Biotechnology , 109 , 1–12. https://doi.org/10.1007/s00253-025-13414-4
- Żbikowska, K., Dolka, B., Kalińska, A., Slósarz, J., Gołębiewski, M., Łoś, M., Strzałkowska, N., Janocha, A., Bień, D., & Michalczuk, M. (2025). In vitro evaluation of the effects of silver nanoparticles on Enterococcus faecalis cells’ viability. Animal Science Papers and Reports , 43 , Article 3. https://doi.org/10.2478/aspr-2025-0022
- Ochirbat, E., Zbonikowski, R., Sulicka, A., Bończak, B., Bonarowska, M., Łoś, M., Malinowska, E., Hołyst, R., & Paczesny, J. (2023). Heteroaggregation of virions and microplastics reduces the number of active bacteriophages in aqueous environments. Journal of Environmental Quality , 52 , Article 3. https://doi.org/10.1002/jeq2.20459
- Raza, S., Folga, M., Łoś, M., Foltynowicz, Z., & Paczesny, J. (2022). The effect of zero-valent iron nanoparticles (nZVI) on bacteriophages. Viruses , 14 , Article 5. https://doi.org/10.3390/v14050867
- Richter, Ł., Stevens, C. A., Silva, P. J., Julià, L. R., Malinverni, C., Wei, L., Łoś, M., & Stellacci, F. (2022). Peptide-grafted nontoxic cyclodextrins and nanoparticles against bacteriophage infections. ACS Nano , 16 , Article 11. https://doi.org/10.1021/acsnano.2c07896
- Richter, Ł., Księżarczyk, K., Paszkowska, K., Janczuk-Richter, M., Niedziółka-Jönsson, J., Gapiński, J., Łoś, M., Hołyst, R., & Paczesny, J. (2021). Adsorption of bacteriophages on polypropylene labware affects the reproducibility of phage research. Scientific Reports , 11 , 1–12. https://doi.org/10.1038/s41598-021-86571-x
- Filipiak, M., Łoś, J. M., & Łoś, M. (2020). Efficiency of induction of Shiga-toxin lambdoid prophages in Escherichia coli due to oxidative and antibiotic stress depends on the combination of prophage and the bacterial strain. Journal of Applied Genetics , 61 , Article 1. https://doi.org/10.1007/s13353-019-00525-8
- López-Laguna, H., L. Sánchez-García, N. Serna, E. Voltà-Durán, J. M. Sánchez, A. Sánchez-Chardi, U. Unzueta, M. Łoś, A. Villaverde and E. Vázquez (2020). “Engineering Protein Nanoparticles Out from Components of the Human Microbiome.” Small 16(30): e2001885.
- Filipiak, M., J. M. Łoś and M. Łoś (2020). “Efficiency of induction of Shiga-toxin lambdoid prophages in Escherichia coli due to oxidative and antibiotic stress depends on the combination of prophage and the bacterial strain.” J Appl Genet 61(1): 131-140.
- Matuła, K., Ł. Richter, M. Janczuk-Richter, W. Nogala, M. Grzeszkowiak, B. Peplińska, S. Jurga, E. Wyroba, S. Suski, H. Bilski, A. Silesian, H. A. R. Bluyssen, N. Derebecka, J. Wesoły, J. M. Łoś, M. Łoś, P. Decewicz, L. Dziewit, J. Paczesny and R. Hołyst (2019). “Phenotypic plasticity of Escherichia coli upon exposure to physical stress induced by ZnO nanorods.” Sci Rep 9(1): 8575.
- Skowron, P. M., A. M. Kropinski, J. Zebrowska, L. Janus, K. Szemiako, E. Czajkowska, N. Maciejewska, M. Skowron, J. Łoś, M. Łoś and A. Zylicz-Stachula (2018). “Sequence, genome organization, annotation and proteomics of the thermophilic, 47.7-kb Geobacillus stearothermophilus bacteriophage TP-84 and its classification in the new Tp84virus genus.” PLoS One 13(4): e0195449.
- Richter, Ł., K. Bielec, A. Leśniewski, M. Łoś, J. Paczesny and R. Hołyst (2017). “Dense Layer of Bacteriophages Ordered in Alternating Electric Field and Immobilized by Surface Chemical Modification as Sensing Element for Bacteria Detection.” ACS Appl Mater Interfaces 9(23): 19622-19629.
- Janczuk, M., Ł. Richter, G. Hoser, J. Kawiak, M. Łoś, J. Niedziółka-Jönsson, J. Paczesny and R. Hołyst (2017). “Bacteriophage-Based Bioconjugates as a Flow Cytometry Probe for Fast Bacteria Detection.” Bioconjug Chem 28(2): 419-425.
- Golec, P., K. Żelechowska, J. Karczewska-Golec, J. Karczewski, A. Leśniewski, M. Łoś, G. Węgrzyn and A. M. Kłonkowski (2017). “Bacteriophages as Factories for Eu(2)O(3) Nanoparticle Synthesis.” Bioconjug Chem 28(7): 1834-1841.
- Żelechowska, K., J. Karczewska-Golec, J. Karczewski, M. Łoś, A. M. Kłonkowski, G. Węgrzyn and P. Golec (2016). “Phage-Directed Synthesis of Photoluminescent Zinc Oxide Nanoparticles under Benign Conditions.” Bioconjug Chem 27(9): 1999-2006.
- Szot-Karpińska, K., P. Golec, A. Leśniewski, B. Pałys, F. Marken, J. Niedziółka-Jönsson, G. Węgrzyn and M. Łoś (2016). “Modified Filamentous Bacteriophage as a Scaffold for Carbon Nanofiber.” Bioconjug Chem 27(12): 2900-2910.
- Krasowska, A., A. Biegalska, D. Augustyniak, M. Łoś, M. Richert and M. Łukaszewicz (2015). “Isolation and Characterization of Phages Infecting Bacillus subtilis.” Biomed Res Int 2015: 179597.
- Lesniewski, A., M. Los, M. Jonsson-Niedziółka, A. Krajewska, K. Szot, J. M. Los and J. Niedziolka-Jonsson (2014). “Antibody modified gold nanoparticles for fast and selective, colorimetric T7 bacteriophage detection.” Bioconjug Chem 25(4): 644-648.
- Kannan, P., M. Los, J. M. Los and J. Niedziolka-Jonsson (2014). “T7 bacteriophage induced changes of gold nanoparticle morphology: biopolymer capped gold nanoparticles as versatile probes for sensitive plasmonic biosensors.” Analyst 139(14): 3563-3571.
- Golec, P., J. Karczewska-Golec, M. Łoś and G. Węgrzyn (2014). “Bacteriophage T4 can produce progeny virions in extremely slowly growing Escherichia coli host: comparison of a mathematical model with the experimental data.” FEMS Microbiol Lett 351(2): 156-161.
- Golec, P., J. Karczewska-Golec, B. Voigt, D. Albrecht, T. Schweder, M. Hecker, G. Węgrzyn and M. Łoś (2013). “Proteomic profiles and kinetics of development of bacteriophage T4 and its rI and rIII mutants in slowly growing Escherichia coli.” J Gen Virol 94(Pt 4): 896-905.
- Bloch, S., B. Nejman-Faleńczyk, J. M. Łoś, S. Barańska, K. Łepek, A. Felczykowska, M. Łoś, G. Węgrzyn and A. Węgrzyn (2013). “Genes from the exo-xis region of λ and Shiga toxin-converting bacteriophages influence lysogenization and prophage induction.” Arch Microbiol 195(10-11): 693-703.
- Łoś, M. and G. Węgrzyn (2012). “Pseudolysogeny.” Adv Virus Res 82: 339-349.
- Los, M. (2012). “Minimization and prevention of phage infections in bioprocesses.” Methods Mol Biol 834: 305-315.
- Łoś, M. (2012). “The Good, the bad and the ugly.” European Biopharmaceutical Review 2012: 68-72
- Loś, J. M., M. Loś, A. Węgrzyn and G. Węgrzyn (2012). “Altruism of Shiga toxin-producing Escherichia coli: recent hypothesis versus experimental results.” Front Cell Infect Microbiol 2: 166.
- Jakubowska-Deredas, M., A. Jurczak-Kurek, M. Richert, M. Łoś, M. Narajczyk and B. Wróbel (2012). “Diversity of tailed phages in Baltic Sea sediment: large number of siphoviruses with extremely long tails.” Res Microbiol 163(4): 292-296.
- Golec, P., J. Karczewska-Golec, M. Loś and G. Węgrzyn (2012). “Novel ZnO-binding peptides obtained by the screening of a phage display peptide library.” J Nanopart Res 14(11): 1218.
- Klawitter, E., D. Kunikowska, E. Tokarska-Pietrzak, H. Dziadziuszko, J. M. Łoś, P. Golec, G. Węgrzyn and M. Łoś (2011). “Specific detection of Salmonella enterica and Escherichia coli strains by using ELISA with bacteriophages as recognition agents.” European Journal of Clinical Microbiology & Infectious Diseases 30, 1067-1073
- Loś, J. M., M. Loś and G. Węgrzyn (2011). “Bacteriophages carrying Shiga toxin genes: genomic variations, detection and potential treatment of pathogenic bacteria.” Future Microbiol 6(8): 909-924.
- Łoś, M. (2011). “Phage in Therapy and Prophylaxis – History and Future Prospects.” International Pharmaceutical Industry. 3, 30-34
- Łoś, M. (2011). “The age of phage.” European Biopharmaceutical Review 56, 58-62
- Golec, P., A. Wiczk, J. M. Łoś, G. Konopa, G. Węgrzyn and M. Łoś (2011). “Persistence of bacteriophage T4 in a starved Escherichia coli culture: evidence for the presence of phage subpopulations.” J Gen Virol 92(Pt 4): 997-1003.
- Golec, P., K. Dąbrowski, M. S. Hejnowicz, A. Gozdek, J. M. Loś, G. Węgrzyn, M. B. Lobocka and M. Loś (2011). “A reliable method for storage of tailed phages.” J Microbiol Methods 84(3): 486-489.
- Los, M. (2010). “Contamination concerns.” European Biopharmaceutical Review, 51, 78-80
- Loś, J. M., M. Loś, A. Wegrzyn and G. Wegrzyn (2010). “Hydrogen peroxide-mediated induction of the Shiga toxin-converting lambdoid prophage ST2-8624 in Escherichia coli O157:H7.” FEMS Immunol Med Microbiol 58(3): 322-329.
- Golec, P., A. Wiczk, A. Majchrzyk, J. M. Łoś, G. Węgrzyn and M. Łoś (2010). “A role for accessory genes rI.-1 and rI.1 in the regulation of lysis inhibition by bacteriophage T4.” Virus Genes 41(3): 459-468.
- Glinkowska, M., J. M. Loś, A. Szambowska, A. Czyz, J. Całkiewicz, A. Herman-Antosiewicz, B. Wróbel, G. Wegrzyn, A. Wegrzyn and M. Loś (2010). “Influence of the Escherichia coli oxyR gene function on lambda prophage maintenance.” Arch Microbiol 192(8): 673-683.
- Nejman, B., J. M. Loś, M. Łoś, G. Wegrzyn and A. Wegrzyn (2009). “Plasmids derived from lambdoid bacteriophages as models for studying replication of mobile genetic elements responsible for the production of Shiga toxins by pathogenic Escherichia coli strains.” J Mol Microbiol Biotechnol 17(4): 211-220.
- Loś, J. M., M. Loś, G. Wegrzyn and A. Wegrzyn (2009). “Differential efficiency of induction of various lambdoid prophages responsible for production of Shiga toxins in response to different induction agents.” Microb Pathog 47(6): 289-298.
- Łoś, M., J. M. Łoś and G. Wegrzyn (2008). “Rapid identification of shiga toxin-producing Escherichia coli (STEC) using electric biochips.” Diagn Mol Pathol 17(3): 179-184.
- Loś, J. M., P. Golec, G. Wegrzyn, A. Wegrzyn and M. Loś (2008). “Simple method for plating Escherichia coli bacteriophages forming very small plaques or no plaques under standard conditions.” Appl Environ Microbiol 74(16): 5113-5120.
- Łoś, M., P. Golec, J. M. Łoś, A. Weglewska-Jurkiewicz, A. Czyz, A. Wegrzyn, G. Wegrzyn and P. Neubauer (2007). “Effective inhibition of lytic development of bacteriophages lambda, P1 and T4 by starvation of their host, Escherichia coli.” BMC Biotechnol 7: 13.
- Los, M. (2006). “Coliphages in their natural environment: strategies of survival.” In Modern Bacteriophage Biology and Biotechnology. Research Signpost, Kerala, India. 103-114.
- Nieradko, J., M. Los (2006). “Strategies by which a bacterial population can survive phage infections.” In Modern Bacteriophage Biology and Biotechnology. Research Signpost, Kerala, India. 115-128
- Los, M. (2006). “Virus detection today.” In Modern Bacteriophage Biology and Biotechnology. Research Signpost, Kerala, India. 129-140
- Neubauer, P., B. Mathiszik, M. Los (2006). “Simulation tool for estimation of optimal bacterial growth rate to reduce effects caused by bacteriophages.” In Modern Bacteriophage Biology and Biotechnology. Research Signpost, Kerala, India. 153-164
- Los, M., J. M. Los, L. Blohm, E. Spillner, T. Grunwald, J. Albers, R. Hintsche, G. Wegrzyn (2005). “Rapid detection of viruses using electrical biochips and anti-virion sera.” Letters in Applied Microbiology, 40, 479-85
- Łos, M., A. Czyz, E. Sell, A. Wegrzyn, P. Neubauer and G. Wegrzyn (2004). “Bacteriophage contamination: is there a simple method to reduce its deleterious effects in laboratory cultures and biotechnological factories?” J Appl Genet 45(1): 111-120.
- Gabig-Ciminska, M., M. Los, A. Holmgren, J. Albers, A. Czyz, R. Hintsche, G. Wegrzyn and S. O. Enfors (2004). “Detection of bacteriophage infection and prophage induction in bacterial cultures by means of electric DNA chips.” Anal Biochem 324(1): 84-91.
- Los, M., G. Wegrzyn and P. Neubauer (2003). “A role for bacteriophage T4 rI gene function in the control of phage development during pseudolysogeny and in slowly growing host cells.” Res Microbiol 154(8): 547-552.
- Gabig, M., A. Herman-Antosiewicz, M. Kwiatkowska, M. Los, M. S. Thomas and G. Wegrzyn (2002). “The cell surface protein Ag43 facilitates phage infection of Escherichia coli in the presence of bile salts and carbohydrates.” Microbiology 148(Pt 5): 1533-1542.
- Czyz, A., M. Los, B. Wrobel, G. Wegrzyn (2001). “Inhibition of spontaneous induction of lambdoid prophages in Escherichia coli cultures: simple procedures with possible biotechnological applications.” BMC Biotechnology, 1, 1